Engineering and Directed Evolution of Proteins
The motivation of our work is to enable novel biomedical applications and therapies through the use of engineered proteins. However, the problems to be solved to reach this goal are typically very basic and fundamental in nature. They often require to investigate the biophysical and structural aspects of proteins, and thus this research also helps us to better understand proteins. Very often, also the development of new technologies is required, such as for directed evolution.
Because of the complex nature of these challenges, our lab is very interdisciplinary. It ranges from protein design and modeling, to directed evolution, to structural biology and biophysical investigations, to cell culture work and animal work. Several projects have gone through all these stages. All aspects are connected by our core ability to engineer proteins — in many ways — and these combined efforts made many projects possible in the first place.
One key capability we have developed over the years is to create novel binding proteins, e.g., the DARPin technology we have invented, and to use them in numerous applications. Another is the ability to create stable protein variants through design and evolution, which has helped in creating crystallizable G protein-coupled receptors, stable antibodies, and stable intracellular binding proteins. An area of particular interest has been the use of engineered proteins in targeted tumor therapies.
Main Research Goals
Protein Engineering Applied to Tumor Targeting
We take a very broad approach to the investigation of tumor targeting, exploiting the protein engineering capabilities we have developed, and to establish rules and principles for designing optimal therapeutics. This also involves the structure and function of the target receptors.
We have been using both our Designed Ankyrin Repeat Proteins (DARPins) and synthetic antibodies in these endeavors. Our high-throughput selection facility gives us access to binders to essentially any target, and through the ability to state-of-the-art modeling, design and directed evolution, we are not limited to existing components, but can generate new ones.
Our focus is on fundamental studies, to understand the interdependencies of affinity, valency, size, stability and effector engineering on in vivo efficacy. This has included many new formats of creating bispecific and multispecific formats, protein-drug conjugates, and new ways of precisely controlling pharmacokinetics.
Another goal is the targeting of specific intracellular proteins and triggering their degradation. We have developed a system for receptor-specific uptake of binding proteins to the cytosol, to achieve two layers of specificity, one for the cell type, one for the target.
Because of the wide range of technologies available in our research group, we can bring projects from biophysical characterization of newly designed proteins, to tests in cell culture, and to measuring biodistribution and efficacy in animal models.
For the use of shielded, retargeted Adenoviruses in tumor targeting, please see the corresponding section.
See our publications, topic Tumor Targeting.
Gene Therapy: Shielded, Retargeted Adenovirus
The goal of this research direction is to develop the technologies for the body to produce its own combination of therapeutic proteins — when and where they are needed. To achieve this, we have developed a gene delivery platform based on special versions of adenovirus.
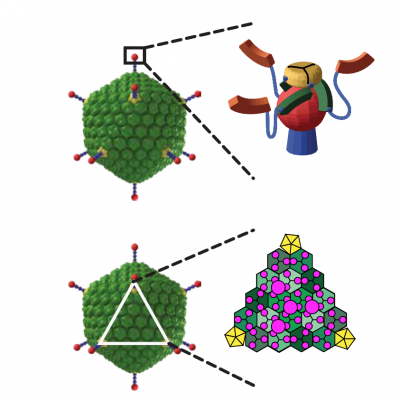
These adenoviral vectors are specific, as they can be programmed to use any cellular surface receptor of choice for uptake and infection. They are safe, as they can have no viral genes at all, cannot proliferate, and their DNA does not integrate into the genome. They have a very large genome (35 kB), and thus can encode multiple genes at the same time. Finally, they are covered by a designed protein shield that hides them from interacting with the immune system and other components of the body.
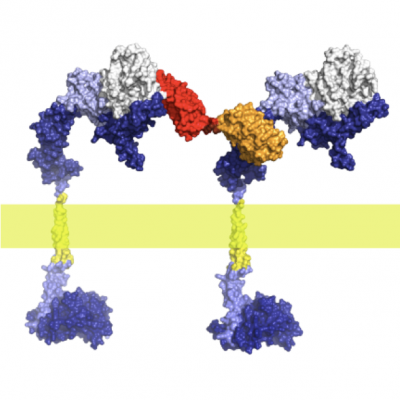
To achieve cell-specific infections, we make use of our protein engineering technologies, especially our DARPin technology of creating binding proteins, with which we can very rapidly create different «adapters» for the same virus. The adapters bind extremely tightly to the virus and thus confer new exchangeable targeting specificities. These adapters have been directed to a range of tumor cells, stromal cells and cells of the immune system, with the portfolio constantly increasing.
See our publications, topic Adenovirus.
G-protein coupled receptors
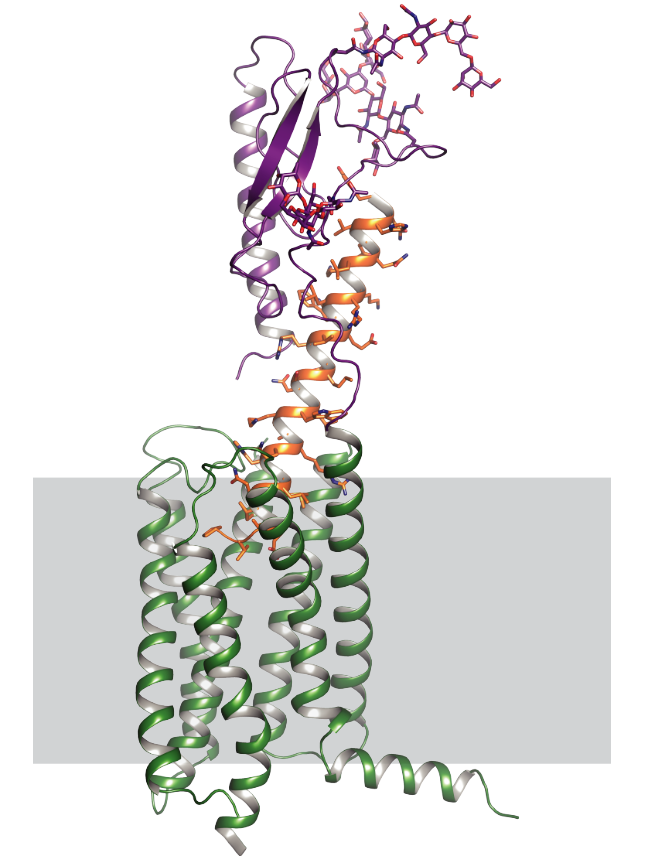
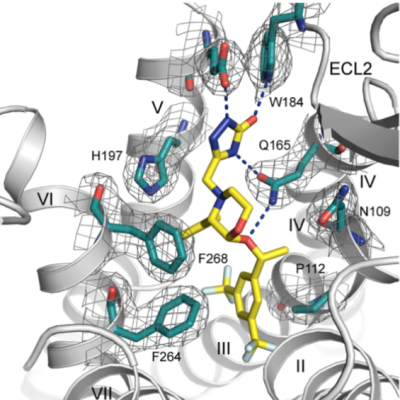
G-protein-coupled receptors regulate a large part of human pharmacology and are the targets of a large fraction of all pharmaceuticals in use today. They undergo characteristic structural changes upon binding antagonists or agonists, and these compounds can be of widely different chemical classes. To understand how specific agonists and antagonists mediate their function, and how to design future active ligands, it is essential to have access to high-resolution structural information. The highest resolution data come from x-ray crystallography, with NMR giving important information on dynamics.
Unfortunately, many wild-type receptors express only in very low yields, are not stable enough to give crystals, or can’t be purified in native form. All of these problems have been solved by a series of protein engineering methods that our laboratory has developed (see Technology Section).
This has allowed us to investigate several GPCRs of different families in structural and functional studies, in different states with different ligands, as well as for experimental screening of ligands and antibodies. Among them were the neurotensin receptor, the parathyroid hormone receptor, the neurokinin receptor, the oxytocin receptor, and several others which are currently being worked on.
See our publications, topic GPCRs.
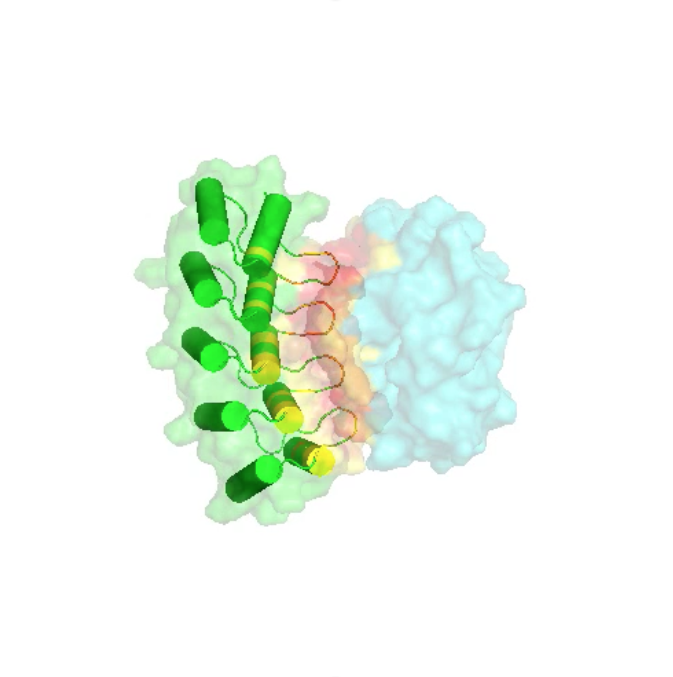
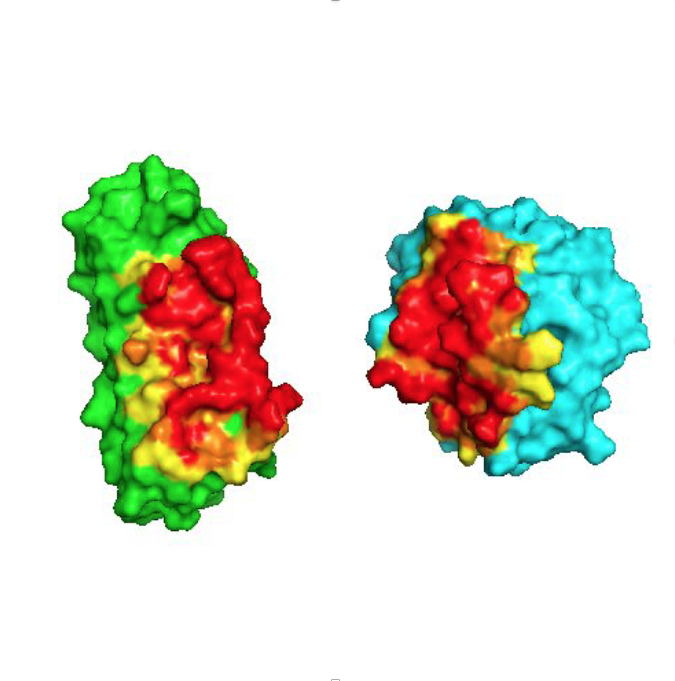
Designed Armadillo Repeat Proteins for creating modular, sequence-specific binding to linear epitopes
One of the fundamental prerequisites for almost all biological and biomedical research are specific, well-performing detection agents. To move beyond the unreliable performance of many commercial monoclonal antibodies, we have been working on a completely orthogonal technology. It is aimed at detecting linear epitopes in a completely modular way, amino acid by amino acid, from pre-selected and pre-designed modules, and in a sequence-specific manner.
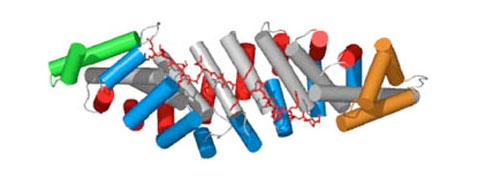
The basis of this technology is synthetic Armadillo Repeat Proteins, which have the ability to bind linear, stretched-out epitopes. A unique feature of this modular binding strategy is that the affinity depends on the length of the peptide and the protein, and thus it can always be adjusted to the «sweet spot» needed for particular applications and experiments.
Using a combination of design, directed evolution and structure determinations by crystallography and NMR, this project brings together all aspects of protein engineering to create the required pockets. These novel pockets have been assembled within Armadillo Repeat Proteins, to confer new sequence-specific binding, and for an ever-increasing number of sequence motifs.
See our publications, topic Armadillo Repeat Proteins.
We develop advanced technologies
Many of the studies in the various projects required technologies that were simply not available at the time. Therefore, we have always been developing novel technologies to be able to greatly expand our toolbox, in order to make certain developments in molecular engineering possible in the first place.
Designed Ankyrin Repeat Proteins as Scaffolds for Selective Binding
Over the last 20 years, we have developed Designed Ankyrin Repeat Proteins (DARPins) as a platform for specific binding proteins. This has meanwhile become a mature technology, which is now also used to develop therapeutics for a variety of diseases. By using consensus design, different synthetic libraries of ankyrin repeat proteins had been created. From these, the desired binders are selected by display technologies such as ribosome display.
Why are DARPins so useful? They are very stable and do not have disulfide bonds and therefore, unlike most antibodies, also work inside the cytosol. They can be prepared in very large amounts from E. coli, are very specific, and often show affinities in the picomolar range. They are obtained from synthetic libraries we have created, using technologies such as ribosome display (in a high-throughput facility), but also using phage display or yeast display.
What can you do with them? They have been used academically in a very wide spectrum of applications, ranging from crystallization chaperones, including for membrane proteins, scaffolds for electron microscopy, as intracellular inhibitors, extracellular receptor ligands, sensors, virus-retargeting adapters, or affinity ligands. Many fusion protein designs would have been difficult to achieve with recombinant antibodies, as DARPins are very resistant to aggregation and misfolding. They have also been used to study fundamental questions of protein folding and design.
See our publications, topic DARPins.
Antibody engineering
Over the years, we have investigated and engineered stable antibody frameworks with significantly better properties than natural ones. This work has been the basis of synthetic antibody libraries, which have given rise to many useful binders, being the basis of therapeutic development. It has also led to a stability-engineered antibody against EpCAM in our lab being now in phase 3 clinical trials. Using the directed evolution technologies (see Ribosome Display), we have evolved extremely tight binders in vitro against proteins, peptides and small molecules, up to 1 pM (a monovalent scFv against the prion protein). Additionally, more fundamental questions have been addressed regarding the in vivo folding and stability, using scFv, Fab, IgG expressed in prokaryotic or eukaryotic cells. This work has helped to elucidate what features prevent misfolding and aggregation.
In recent years, we have used these capabilities to generate therapeutic IgGs, bispecific and multispecific antibodies of many formats and of different isotypes. Some of these constructs are on a trajectory to clinical trials. Moreover, these capabilities have been used to express IgGs in situ, using the Shielded Retargeted Adenovirus platform.
See our publications, topic Antibody Engineering.
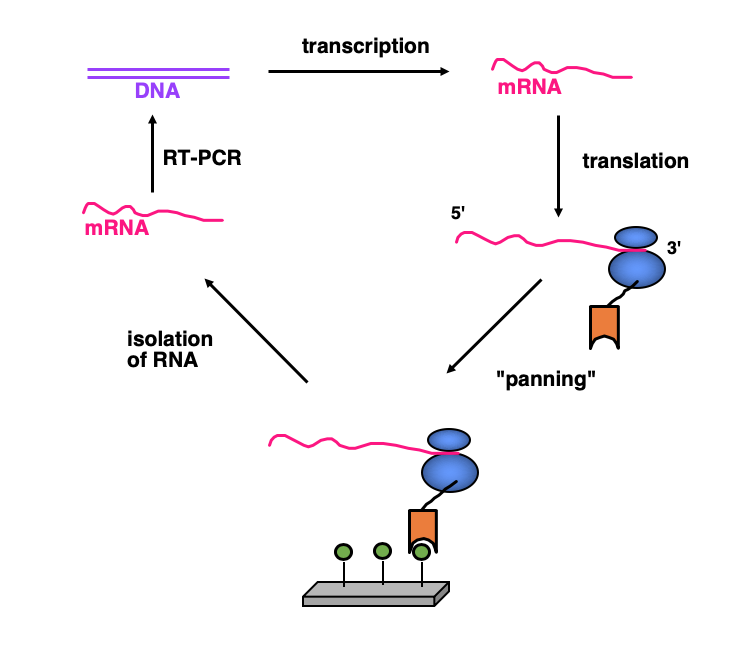
Directed Evolution: Ribosome Display
Ribosome display is a method of cell-free selection and evolution from very large libraries that we have devised and that is now widely used. Since there is no transformation of cells involved, much larger libraries can be screened than with other technologies. Ribosome display has affinity maturation built in, as the errors of a PCR can be exploited, or error-prone PCR can be used to increase the error rate. Ribosome is thus a true in-vitro Darwinian evolution.
Ribosome display has been adapted to a high-throughput mode, where 96 different selections can be done in parallel. It has also been adapted to affinity maturation of individual clones (reaching a monovalent dissociation constant of 1 pM), improvement of protein stability.
See our publications, topic Ribosome Display.
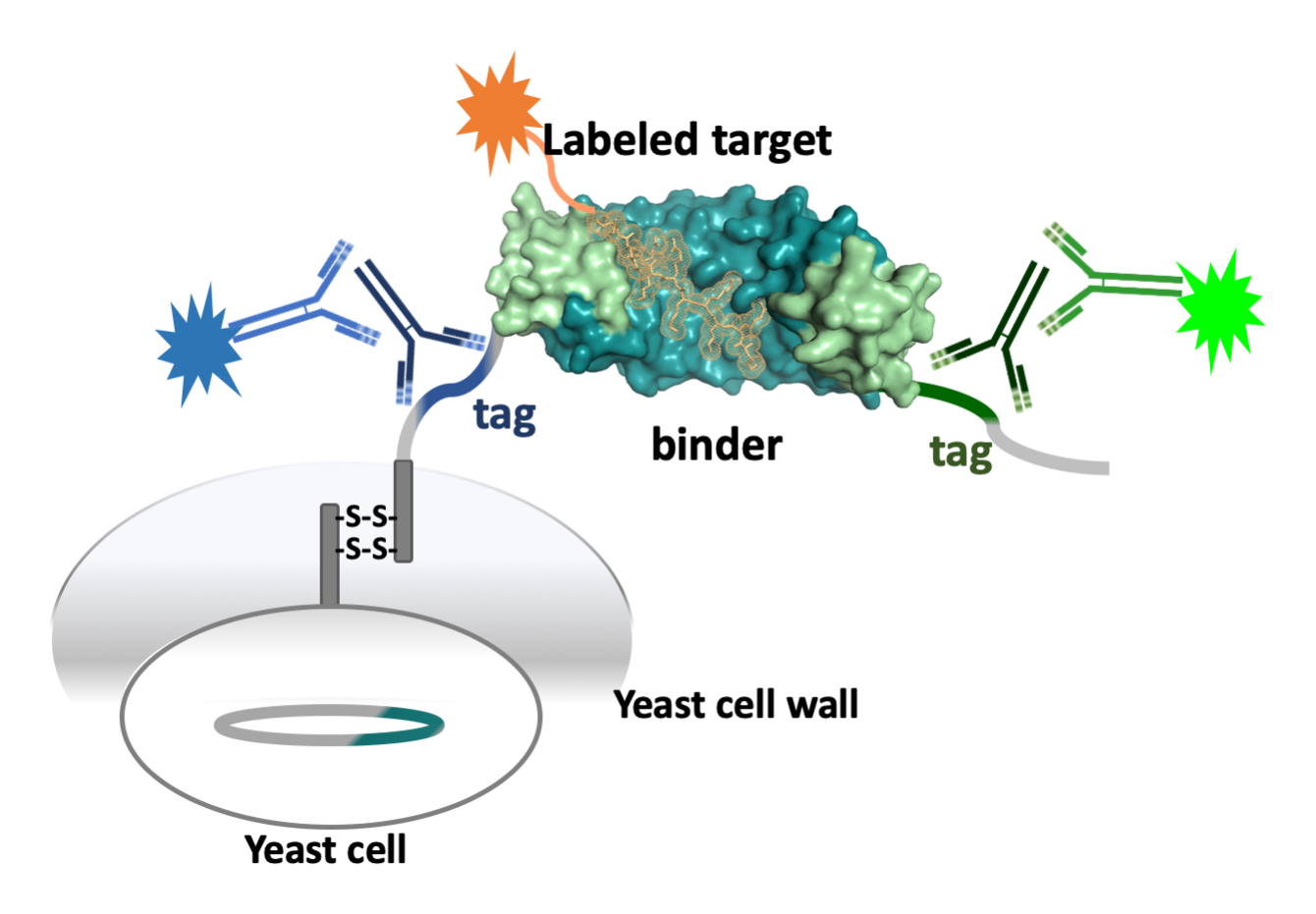
Directed Evolution: Yeast Display
We have adapted yeast display, mostly in the project on Armadillo Repeat Proteins. A new strategy in using yeast display has been devised, focusing mostly on specificity. It succeeds in competing off binders which predominantly recognize other targets (unspecific binders), but, even more importantly, also removes unselective binders, which, in addition to the correct target, also recognize related targets. Yeast display is using FACS, and the art of selection and counterselection has been pivotal in obtaining high-specificity binders with Armadillo Repeat Proteins.
See our publications, topic Other Selection Methods.
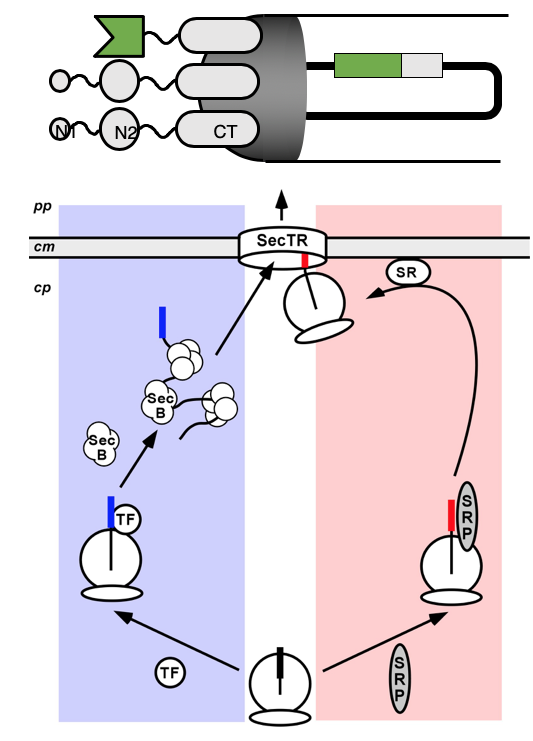
Directed Evolution: Phage Display and Related Methods
Phage display has been adapted in our group to make it compatible with a wider spectrum of proteins to be displayed, by changing the export pathway. This has particularly made DARPins amenable to phage display. Phage display has also been used in our group to select for properties other than mere binding, such as protein stability and enzymatic turnover, as well as coupling it to the infection process.
See our publications, topic Phage Display and Other Selection Methods.
Directed Evolution and Engineering for Stability of GPCRs
One of the biggest challenges in the study of the structure and biophysics of eukaryotic membrane proteins is that these proteins are extremely difficult to handle: they are usually hard to produce in large amounts, they are very unstable, and they often denature when solubilized in detergent. Our lab has devised several technologies to solve these problems. By expressing mutant libraries of the GPCRs in E. coli and yeast, and by supplying fluorescently labeled agonist or antagonist, those variants can be enriched which express more functional receptor. It has turned out that the variants expressing better in E. coli or yeast also express better in baculovirus-infected insect cells and mammalian cells, and are more stable after solubilization in detergent. Additionally, we have developed a directed evolution technology termed CHESS, in which bacteria are individually wrapped in a polymer, and the inside is completely dissolved by detergent, and thus detergent-stable variants of GPCRs can be selected directly using FACS.
Furthermore, very efficient ways of creating and testing hundreds of GPCR mutants for stability have been devised that have been pivotal in obtaining the recent GPCR structures from our laboratory. Finally, new crystallization chaperones have been created that have ultimately allowed structure determination of a number of recalcitrant receptors. All these efforts have not only made eukaryotic membrane proteins amenable to study, but are helping to elucidate the structural rules of stable membrane proteins.
See our publications, topic GPCRs and Other Selection Methods.
Structural biology
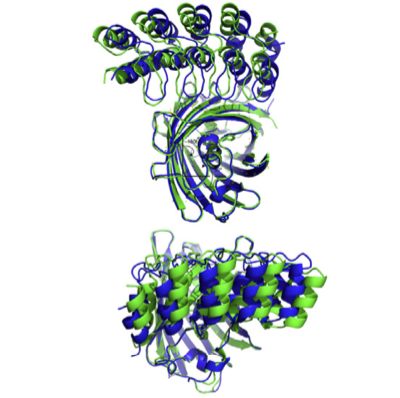
X-ray crystallography has played a pivotal role in all projects. In the GPCR projects, it has allowed to describe the interaction with ligands (agonists and antagonists) with the receptor, as well as the induced conformational changes. In projects with DARPins, it has clarified the binding epitope, and thereby elucidated the mode of action of pharmacologically active DARPins. In projects involving Armadillo Repeat Proteins, it has been part of the protein design cycle, to verify the molecular structure to exact details, and the binding mode of the ligand, to influence the next design cycle.
Conversely, protein engineering has also helped crystallography in general: Using rigidly fused DARPins as new crystallization chaperones, several structures could be determined that did not give rise to diffracting crystals otherwise.
Cryo-Electron microscopy of surface receptors is being used, both as single particles and in whole cell tomography, in collaboration with the group of Ohad Medalia. Here, the capability of creating tailor made binding proteins is playing a key role. NMR is used in several collaborations, involving all of the main research directions.
See our publications, topic Structures.
Protein Design and Molecular Modeling
Protein Design and Molecular Modeling has played a role in almost all applications. Convenient tools and scripts have been built for homology modeling, especially for antibodies, but also for rigid fusion proteins. Rosetta has been used extensively for docking, and evaluating mutations for stability. All libraries had a component of consensus design built in.
Small molecule docking has been carried out in collaboration with other groups